PCR and qPCR Enzymes and Reagents
FailSafe PCR PreMix Selection Kit
PCR optimisation kit for end-point applications. Includes everything needed to optimise PCR performance and achieve consistent, high-fidelity results for every DNA template…
Before purchasing, we strongly recommend first optimising your PCR conditions using the FailSafe PCR PreMix Selection Kit in order to know which 2X PreMixes to choose with your PCR System kit.
Includes FailSafe PCR Enzyme Mix, a blend of thermostable enzymes containing a 3′→5′ proofreading enzyme for high fidelity, plus your choice of 1, 2, or 8 of the FailSafe PCR 2X PreMixes (100 uL each). 2X PreMixes contain dNTPs, buffer, different concentrations of MgCl2, and PCR Enhancer with betaine.
PCR System kit for end-point applications. Includes everything needed for successful PCR performance. Achieve consistent, high-fidelity results for every DNA template.
PCR and qPCR Enzymes and Reagents
PCR optimisation kit for end-point applications. Includes everything needed to optimise PCR performance and achieve consistent, high-fidelity results for every DNA template…
PCR and qPCR Enzymes and Reagents
PCR Enzyme Mix, individual component of the FailSafe PreMix Selection Kit for PCR optimization, and the bundled post - optimization FailSafe PCR System.
PCR and qPCR Enzymes and Reagents
Twelve 2X PreMixes, individual components of the FailSafe PreMix Selection Kit for PCR optimization, and the bundled post - optimization FailSafe PCR System.
The FailSafe PCR System is the post optimization, bundled, and affordable kit that contains everything you need for successful PCR performance. The kit is multiplex PCR compatible, can amplify even the most difficult high-GC templates, and targets up to 20 kb in length.
The PCR system can be purchased after PCR conditions are optimised with the FailSafe PCR PreMix Selection kit, so you know which 2x Premix to choose for your template/primer combination.
Choose between 100, 250, or 1000 Enzyme Mix units, then combine with your top choice of 1, 2, or 8 out of the 12 PCR 2x PreMixes. The Enzyme Mix included has a fidelity of at least three times higher than Taq DNA polymerase.
The FailSafe 2x PreMixes and Enzyme Mix can be purchased with the PCR PreMix Selection Kit for initial optimization, with the PCR System, or, as individual components.
Applications
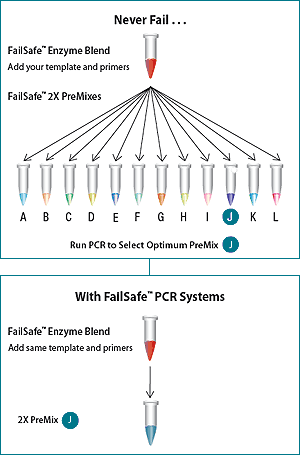
How It Works
Step 1
Start with the FailSafe PCR PreMix Selection Kit to optimise end-point PCR conditions.
Step 2
Analyse and determine which FailSafe PCR 2x PreMixes produce high fidelity results, based on the template/primer pair combination.
Step 3
Purchase the post optimization FailSafe PCR System, which includes the Enzyme Mix plus your choice of 1, 2, or 8 out of the 12 PCR 2x PreMixes, OR, purchase the FailSafe PreMix and Enzyme components individually; and achieve reliable and consistent high-fidelity PCR results.
For a complete explanation of how FailSafe technology works, please read the manual.
 |
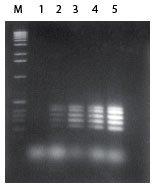 |
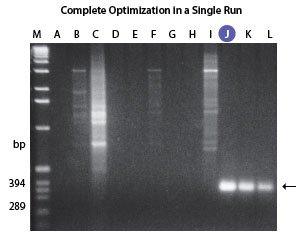
| Figure 1. Amplification of an 80%-85% GC-rich region of the human fragile X gene. PCR was performed using the FailSafe PCR System, schematically depicted above. Lanes A-L show the amplification products resulting from PCR using the 12 FailSafe PCR PreMixes. Lane M, molecular weight marker. In this experiment, optimal amplification was obtained with FailSafe PCR PreMix J. The size of the expected amplicon is indicated by an arrow. | Figure 2. Multiplex PCR of the human CFTR gene. The FailSafe PCR System amplified all five exons of the CFTR gene from as little as 1 ng of human genomic DNA. |
PCR and qPCR Master Mixes
PCR and qPCR Master Mixes
PCR and qPCR Master Mixes
ROX Level
ROX Level
Need some support with placing an order, setting up an account, or finding the right protocol?
Contact us| Europe, Middle East, and Africa | |
|---|---|
| UK | +44 1992 470 757 |
| Germany | +49 30 5304 2200 |
| North America, Latin America | |
| Wisconsin, USA | +1 888 575 9695 |
| Asia Pacific | |
| China | +8621-22509000 |
| Singapore | +65 6734 4800 |